Trigger warning¶
When Dr. Capehart does this in a live session, you get to hear the full (and sometimes uncensored) rant about when a certain PhD student brought their child to sit (in his good chair) with a very distinctive cough to find out by an email the day after that he had been exposed to Whopping Cough.
Get your Tdap!
Overall Goals and Objectives¶
The goal here is to demonstrate using a Jupyter Notebook to present a fully integrated project combining code, figures, and narration. In this example, we are presenting a classic Epidemic Model.
In this exercise, our objectives will be:
- Using the markdown features of the Jupyter Notebook to create a report on exploring these equations
- Coding the K-McK equations in Python and graphically presenting the output.
- Exploring the relationship between the Basic Reproductive Ratio (The "R₀") and Herd Immunity
- Compare different levels of community immunity (i.e., an inoculation program) on the spread of an outbreak.
Background¶
This Jupyter Notebook presents a simple variant of the classic "Kermack-McKendrick" System. As the name suggests, this code was developed by a pair of British Scientists, William Kermack & Anderson McKendrick. They took previous work and condensed it down into what we now know as the classical set of equations that we know today. It is also often referred to as a "SIR" model named after the three primary populations ("Susceptable," "Infected," and "Recovered"). The paper is available in its original form here. Pre-COVID, we used to have a fourth category, "Fatalities," which under the circumstances are now in bad taste since we are not monsters and use Salk.
Kermack, W.O. and McKendrick A.G. 1927: A contribution to the mathematical theory of epidemics, Proceedings of the Royal Society, A: Mathematical Physical & Engineering Sciences, 115, 700–721, doi:10.1098/rspa.1927.0118.
The modeling system can be used to teach basic principles of modeling and also can be featured in Differential Equation classes. In recent years (pre-COVID) it's also become a resurgence in modeling and Diff-e-Fu™, following the "controversy" over the MMR (Measles-Mumps-Rubella) vaccine and it's debunked relationship with autism*.
*Spoilers: there is no connection. If you want a deep-dive on the "Wakefield MMR Controversy" and an uncomfortable teachable moment on ethics (or lack of) in STEM, Brian Deer has probably one of the best documentaries on the topic, from Wakefield's competing Measles vaccine to his collaboration with someone who thought he could cure autism with his bone marrow (no really!), to the unethical, brutal and unnecessary tests on child test subjects. (My teenage son will point you to a funnier deep-dive based on Deer's work by a certain popular YouTube gamer if you ask him. "I, however, cannot possibly comment.")
Sketching the Model Environment and the Kermack-McKendrick Equations¶
It's always nice to sketch and doodle our models. The black letters will be our variables. (We're not doing the "cordon sanitaire" option (where healthy people wall themselves off from the sick people) in this example.
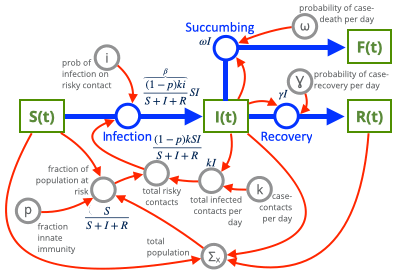
This creates the following system of equations with the labeled parameters above.
$$ \begin{aligned} \frac{dS}{dt} &= -\beta SI \\ \\ \frac{dI}{dt} &= +\beta SI - \gamma I - \omega I \\ \\ \frac{dR}{dt} &= +\gamma I \\ \\ \frac{dF}{dt} &= +\omega I \\ \\ \end{aligned} $$
The above $\beta$ term is a combination of $\kappa$, $p$, and $i$. We will show how they are constructed as we move through this exercise. The above system can be represented by the aforementioned Kermack-McKendrick Equations.
$$\begin{align} \frac{dS}{dt}&=-\beta S I \\ \frac{dI}{dt}&=\beta S I - \gamma I \\ \text{where }\beta &= \frac{i k}{S+I+R} \end{align}$$
The source code and table below list its component variables and their representations. (As your code grows, so should the table!)
| Symbol | Description | Code |
|---|---|---|
| $S(t)$ | Susceptible Population | S[t]; S_long[t] |
| $I(t)$ | Infected Population | I[t]; I_long[t] |
| FIX | FIX | R[t]; R_long[t] |
| Total Impacted Cases | Impacted_Cases | |
| $\beta$ | Net Infection Rate | beta |
| $\gamma$ | Recovery Rate | gamma |
| $i$ | Infection Rate | prob_infect |
| $\kappa$ | Daily Infected Contacts Per Case | kappa |
| $R_0$ | Basic Reproduction Number (The R-naught) | Ro |
| $p_c$ | Threshold Herd Immunity | p_c |
| $p$ | Preexisting Herd Immunity | preexisting_immunity |
| Start Time (Days) | time_start | |
| End Time (Days) | time_end | |
| $\Delta t$ | Time Step (Days) | time_step |
| $t$ | Simulation Time | time; time_long |
| $N$ | Number of Time Steps | N; nt; nt_long |
| Time Iteration Index | t | |
| Number of Immunity Ensembles | n_scenarios | |
| Ensemble Loop Index | p | |
| A Priori Immunity in an Ensemble Loop Index | p | |
| A Priori Immunity in an Ensemble Loop Index | immunity_array[p] | |
| Total Number of Cases in an Ensemble Member | total_cases_long[p] | |
| Peak Number of Infected Cases in an Ensemble Member | max_number_of_cases[p] | |
| Time of Peak Infected Cases in an Ensemble Member | time_of_max_cases[p] |
Libraries¶
Let's start with the libraries
- NumPy for some basic math functions
- Matplotlib for plotting
also...
- IPython for document support (this is just to import a youtube video)
################################################
#
# Libraries
#
import numpy as np
import matplotlib.pyplot as plt
# Notice here that this is using a very different
# method to pull in functions.
#
# I normally don't import libraries this way
#
# This is simply (as the name implies) to bring
# in a youtube video.
from IPython.display import YouTubeVideo
#
################################################
Setting up the Code¶
To do this we will break this process down into several sub-sections
- set up the time extent of our simulation, "Time Control"
- set up the initial state of our players, "Initial or Boundary Conditions"
- set up the rules for the simulation, "Initialize Parameters"
First let’s set up the time control of our simulation. There are variants of this system where you can do truly horrific things to the players to this system such as lose their immunity after a while, add new babies who can get sick easily or have temporary maternal immunity, recovered populations become sterile, or weaker with a higher natural death rate. AES/CEE 615 (Environmental Systems Modeling) explores some of these scenarios at different levels depending on how ~disturbed~ creative the students are for that given semester.
But in this case we are playing a short-game lasting about a month. We’d like some high temporal resolution. So let’s start a simulation of 30 days with a time granularity (time step, $\Delta t$) of about 0.125 days (3-hrs).
################################################
#
# Time Control
#
#
# Total Duration and Time Interval
#
TimePeriod = 30 # days
DeltaT = 3./24. # hours -> days
#
# Number of timesteps from 0 to TimePeriod
#
nTime = int(TimePeriod / DeltaT + 1)
#
# Time Array from 0 to 30
#
# This is different from some array generators
# in python since linspace acutally gives you
# the stopping value you *asked* for! (No
# need to add an extra time step.)
#
Time = np.linspace(start = 0,
stop = TimePeriod, # inclusive must be an integer
num = nTime)
#
################################################
Setting up Initial/Boundary Conditions¶
The above equations need a starting point. In this case, it's our cast of characters.
- Starting Susceptables, $S_0$ (in this run this includes people who are immunized, so technically this just all healthy people).
- Initial Infected Cases, $I_0$ (This/These are our "Patient(s) Zero. Pro-Tip: Don't be these guys.) Immunization Rant will be down further in this notebook!
- Recovered, $R_0$ (not to be confused with another $R_o$ that we will discussed shortly)
- Fatalites, $F_0$ (because as mentioned above, we are monsters.)
################################################
#
# Initial Conditions
#
S_0 = 998.0 # Uninfected People (includes Vaccinated)
I_0 = 1.0 # Patients Zero: Arg! We *HATE* that guy!
R_0 = 0.0 # Recovered: 0 since we're just starting
F_0 = 0.0 # Fatalities: Again, 0 since we're just starting
#
################################################
Creating our Arrays.¶
Here we are going borrow from a certain British TV cooking presenter and repeat our mantra for this class, "maximum satisfaction with minimal effort."
We already have an array as long as we want, our Time[] array. All we have to do is just borrow it, multuply it by zero and we have our arrays for our 4 population groups.
################################################
#
# Setting up our arrays
#
S = Time*0 # Uninfected People (multiplying time by zero)
I = Time*0 # Infected (multiplying time by zero)
R = Time*0 # Recovered (multiplying time by zero)
F = Time*0 # Fatalities (multiplying time by zero)
#
# And we can initialize our first (or zero'th since it's python)
# time step from our above initial conditions
#
k = 0
S[k] = S_0 # Uninfected People (includes Vaccinated)
I[k] = I_0 # Infected
R[k] = R_0 # Recovered
F[k] = F_0 # Recovered
print(" Time[", k, "]=", Time[k],
" S[", k, "]=", S[k],
" I[", k, "]=", I[k],
" R[", k, "]=", R[k],
" F[", k, "]=", F[k])
#
################################################
Time[ 0 ]= 0.0 S[ 0 ]= 998.0 I[ 0 ]= 1.0 R[ 0 ]= 0.0 F[ 0 ]= 0.0
Setting up Basic Disease ~Rules~ Parameters¶
Every model needs a set of quantified rules. For diseases this is difficult to explicitly nail down. As such, we often rely on a set of expected probabilities. Here are the traits we'll be working with
Now onto the disease: We will need to have the
contact rate: the likely number of people who an infected person comes into contact per unit time ($\kappa$)
- Units: probability of infection per risky contact
infection rate: the probability of infection when a truly vulerable person is exposed per unit time ($\kappa$)
- Units: person-to-person contacts (of $S$'s, $I$'s and $R$'s) per sick person per unit time
recovery rate: the probability of recovery per unit time for a given infected person ($\gamma$)
- Units: probability of a given sick person recovering per unit time
fatality rate: the probability per unit time of an infected person dying from the disesase ($\omega$)
- Units: probability of a given sick person to take the eternal dirt nap it per unit time.
$\beta$ from the above equation system is a little more complex than the original case and is a combination of several of these terms. We'll address this later.
- Units: new infections per contact between sick and susceptable people per unit time
immunization fraction: If we want to include factors like artificial or natural immunity ($p$) we can include that, too. Again, we aren't ready for that yet but we'll assume it's zero for now.
- Units: Fraction of people ~smart enough~ not stupid enough to listen to Andrew Wakefield and Jenny McCarthy.
- Education Outreach: my vaccinated child doesn't have autism, he just doesn't like you.
We can start with the following values
| Symbol | Name | Value |
|---|---|---|
| $i$ | Probability of infection | 0.25 |
| $\kappa$ | Sick people contacts w/ other active players | 6.00 |
| $\gamma$ | Probability of Recovery | 0.50 |
You can get this through some unit analysis. If you follow the above diagram you can see how the parts come together. Beta is officially a time dependant field but it is calculated as a function of values form a single time step. This makes this a "diagnostic" variable. $S\left(t\right)$, $I\left(t\right)$, $R\left(t\right)$, and $F\left(t\right)$ since they are represented in the above equations as time-derivatives.
$$\beta\left(t\right)=\frac{i \kappa (1-p)}{S\left(t\right)+I\left(t\right)+R\left(t\right)}$$
################################################
#
# Disease Traits
#
contact_rate = 6.00 # typical number of contacts (exposure) per case per day
infection_rate = 0.25 # probabilty of infection on exposure per case per day
recovery_rate = 0.50 # mean chance of recovery per day per case per day
fatality_rate = 0.10 # mean chance of fatality per case per day
#
# Public Health Traits
#
immunization_fraction = 0.0 # fraction of artificial immunity
#
################################################
Value-added Parameters¶
Basic Reproductioin Number, $R_o$¶
If you've seen the medical thriller "Contagion" the CDC Epidemiologist played by Kate Winslet gives a description of something called the "Basic Reproductive Number," or "R-nought" ($R_o$), factor while plague-splaining the spread of the disease to city planners.
[
# embed youtube video
YouTubeVideo('RGj5blR6IHg')
The $R_o$ describes how many people one infected person will infect.
Typical values for diseases we may encounter in these scenarios include
| Disease | Transmission | R_0 |
|---|---|---|
| Measles | Aerosol | 12–18 |
| Mumps | Respiratory droplets | 10–12 |
| Chickenpox (varicella) | Aerosol | 10–12 |
| COVID-19 (Omicron variant) | Respiratory droplets and aerosol | 9.5 |
| Rubella | Respiratory droplets | 6–7 |
| Polio | Fecal–oral route | 5–7 |
| Pertussis | Respiratory droplets | 5.5 |
| COVID-19 (Delta variant) | Respiratory droplets and aerosol | 5.1 |
| COVID-19 (Alpha variant) | Respiratory droplets and aerosol | 4–5 |
| Conjunctivitis | Dirty Lab Googles! | ~4 |
| "MEV-1" (disease from Contagion) | Hanging out with Gweneth Paltrow (and fomites) | 4 |
| Smallpox | Respiratory droplets | 3.5–6.0 |
| HIV/AIDS | Body fluids | 2–5 |
| SARS | Respiratory droplets | 2–4 |
| Common cold (e.g., rhinovirus) | Respiratory droplets | 2–3 |
| COVID-19 (ancestral strain) | Respiratory droplets and aerosol | 2.9 (2.4–3.4) |
| Diphtheria | Saliva | 2.6 (1.7–4.3) |
| Monkeypox | Physical contact, body fluids, respiratory droplets | 2.1 (1.5–2.7) |
| Influenza (1918 pandemic strain) | Respiratory droplets | 2 |
| Ebola (2014 outbreak) | Body fluids | 1.8 (1.4–1.8) |
| Influenza (2009 pandemic strain) | Respiratory droplets | 1.6 (1.3–2.0) |
| Influenza (seasonal strains) | Respiratory droplets | 1.3 (1.2–1.4) |
| Andes hantavirus | Respiratory droplets and body fluids | 1.2 (0.8–1.6) |
| MERS | Respiratory droplets | 0.5 (0.3–0.8) |
| Nipah virus | Body fluids | 0.5 |
$R_o$'s greater than 1 indicate a disease that will spread through the population. The higher the $R_o$, the faster the spread. $R_o$'s below 1 will likely self-contain due to low probability of infection (not every case will result in a new case, but an influx of sick people may create a critical mass of "players" for some "Susceptables" to lose to the odds). Using our parameters, we can derive $R_o$ via unit analysis as
$$R_o =\frac{i \kappa}{\gamma +\omega}$$
Or if you REALLY want to play with equations in Markdown:
Assumung again that $R=0$ and $S\gg I$...
$$\begin{align*} \frac{dI}{dt}&= \beta S I - \gamma I \\ 0 &= \beta S I - \gamma I \\ 0 &= I \left (\beta S - \gamma \right ) \\ 0 &= \beta S - \gamma \\ \gamma &= \beta S \\ \gamma &= \left( \frac{i k}{S+I+R} \right )S \\ \gamma &= \frac{i k}{S} S \\ \gamma &= i \kappa \\ 1 &= \frac{i k}{\gamma} \\ \text{let } R_0 &= \frac{i k}{\gamma} \\ \text{if you add fatalities: } R_0 &=\frac{i \kappa}{\gamma +\omega} \end{align*}$$
################################################
#
# Basic Reproductive Number
#
Ro = infection_rate * contact_rate / (recovery_rate+fatality_rate)
print("Basic Reproductive Number, Ro =", Ro)
#
################################################
Basic Reproductive Number, Ro = 2.5
#We'reDoingShots!¶
In this case we can play with a fraction of the population that enter the game with either natural immunity or have been immunized. This is the $p$ parameter above.
This also allows us to bring up the idea of herd immunity which is one of the major battlegrounds in the vaccine "controversy."
Let's consider
Assumung that $R=0$ and $S\gg I$,
$$\begin{aligned} \frac{dI}{dt} &=\left[ \beta = \frac{ i k \left( 1-p \right) }{S+ I+R}\right ] S I - \gamma I + \omega I \\ \frac{dI}{dt} &= \frac{ i k \left[ 1-(p=0) \right] }{S+ \left( I+R\approx 0 \right)} S I - \gamma I + \omega I \\ \frac{dI}{dt} &= \frac{ i k }{S} S I - \gamma I + \omega \\ 0 =i k I - \gamma I + \omega I \\ 0 = \left ( i k - \gamma + \omega \right) I \\ 0 &= i \kappa \left( 1-p \right)- \left( \gamma +\omega \right) \\ i \kappa\left(1-p\right) &= \gamma +\omega \\ 1-p &= \frac{\gamma +\omega }{i \kappa} \\ 1-p &= \frac{1}{R_o} \\ p &= 1 - \frac{1}{R_o} \end{aligned} $$
This "critical" value of public immunity fraction, $p$, or $p_c$, is the "herd immunity." Thus for a $R_o$ of 12 (the low end of the communicability of Measles), to force the functional $R_o$ for Measles down to zero or less, over 90% of the populus must have artificial immunity.
################################################
#
# Herd Immunity
#
herd_immunity = 1 - 1/Ro
print("Herd Immunity Threshold, pc =",
herd_immunity)
#
################################################
Herd Immunity Threshold, pc = 0.6
Drafting our Prognostic and Diagnostic Equations¶
In this scenario we are going to show off SciPy's integrating features. Most of them will fit into the following setup.
We will need a single function that contains all of our prognostic equations, $\frac{dS}{dt}$, $\frac{dI}{dt}$, $\frac{dR}{dt}$, and $\frac{dF}{dt}$. Since two of our equations need a value for $\beta$, that gets put in there too.
The Functions for our finite differencing approach¶
Here we will create a single function that bundles all of our equations. In the Differential Equations Hardcore Version of this exercise, we will use a single function with all four equations.
Also we will have an equation for $\beta$ since we will have an opportunity to get fancy with that function as we play.
################################################################################################
#
# Our Four Beta, S, I, R, F function
#
# We pass a vector of [S, I, R, F] through our system.
#
#
# The Beta Function (the rate of aquisition of disease per
# susceptable-infected contacts.)
#
def beta(S,I,R):
beta_out = (1-immunization_fraction)*(infection_rate*contact_rate)/(S+I+R)
return(beta_out)
#
# The Susceptable Equation
#
def dSdt(S,I,R):
dSdt_out = -beta(S,I,R) * I * S
return(dSdt_out)
#
# The Infected Equation
#
def dIdt(S,I,R):
dIdt_out = beta(S,I,R) * I * S - (recovery_rate + fatality_rate) * I
return(dIdt_out)
#
# The Recovered Equation
#
def dRdt(I):
dRdt_out = recovery_rate * I
return(dRdt_out)
#
# The Fatalities Equation
#
def dFdt(I):
dFdt_out = fatality_rate * I
return(dFdt_out)
#
################################################
Time to Bunny-Hop through Time! Euler's Method¶
We are doing a finite differencing approach similar to our earlier adventures where we learned programming with Mathcad.
There are fancier ways to do it but the goal here is more to show you some traditional structures in programming, such as loops and if-then-blocks.
Here is the basic math of any generic prognostic (time-dependant) function.
- Take a differential equations that's a function of time. That $f_x(x)$ function represents all the forcings that are being inflicted on our mystery value x that change its value over time. $$\frac{dx}{dt}=f(x)$$
- Whistle in the dark through your Calc 1 class and hope that $f_x(x)$ doesn't change much over a small discrete change of your independant integrating value (for us that's time), a.k.a., $\Delta t$ $$\frac{\Delta x}{\Delta t}=f(x)$$
- Let's now treat time like a numberline with distinct intervals of space one $\Delta t$ with our understanding of what's going on only resolvable at those discrete points on the hopscotch board that is now our modeled universe. To move forward we base our change on $x$ on our jump to the current time-step for which we're calculating, $t_{\tau}$, on our values that we just solved on our previous timestep, $t_{\tau-1}$. (Hence it's important to chose timesteps where our valus of x don't change MUCH as we move in our steps forward in time. This often means small values of $\Delta t$. Much as in our scripts were we played with Pi using Reimann Sums)
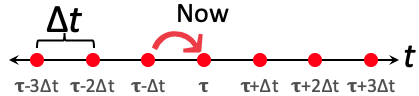
$$\begin{aligned} \frac{\Delta x}{\Delta t}=\frac{x_{\tau}-x_{\tau-1}}{\Delta t=t_{\tau}-t_{\tau-1}}&=f_x(x_{\tau-1})\\ \frac{x_{\tau}-x_{\tau-1}}{\Delta t}&=f_x(x_{\tau-1}) \end{aligned}$$
- And from there it's just a little Algebra-Fu™️ to solve for the value of x on our landing point (a.k.a., $x_{\tau}$)
$$\begin{aligned} \frac{x_{\tau}-x_{\tau-1}}{\Delta t}&=f_x(x_{\tau-1}) \\ x_{\tau}-x_{\tau-1}&=f_x(x_{\tau-1})\Delta t \\ x_{\tau} &= x_{\tau-1}+f_x(x_{\tau-1})\Delta t \end{aligned}$$
Applying this to our scenario we have four equations moving together through time.
$$\begin{aligned} S_{\tau} &= S_{\tau-1}+f_S(x_{\tau-1})\Delta t \\ I_{\tau} &= I_{\tau-1}+f_I(x_{\tau-1})\Delta t \\ R_{\tau} &= R_{\tau-1}+f_R(x_{\tau-1})\Delta t \\ F_{\tau} &= F_{\tau-1}+f_F(x_{\tau-1})\Delta t \end{aligned}$$
Our $f_S$, $f_I$, $f_R$, and $f_F$ are the functions we created above!
Overall this method is called finite differencing. The specific method of marching forward-in-time shown above is call Euler's Method.
To solve this we just need to put things into a loop!
For demonstration purposes, we're going to set this up one step at a time but copying our previous work each step where we add a feature or two to show how we do it specifically in the Python language.
Just remember that above we already have the following "victory laps" already done:
- Created our Time and Time Step Variables
- Created our Modeling Variable Arrays & Initial Conditions
- Created our Model Parameters (e.g., infection rate, recovery rate...)
- Created our Functions for the Kermack-McKendrick Equations.
So onward...
Victory Lap #5. Build the time loop.¶
This is going to be a very simple loop that walks through our time. Here we can loop through our time indices which we'll call $k$ which will go from 0 to nTime.
Remember though that we have to do that wierd thing that Python does where the last falue requested in an array or series is the one AFTER the last one. This is supposedly because this makes it prettier in languages that start with zero (though those of us who have worked with languages that start their indexing at 0 are used to just subtracting one to the number of values you want...).
We'll use numpy.arange() for this
################################################
#
# Step 5: Build the Time Loop
# We are also going to
#
k = 0
print(" Time[", k, "]=", Time[k])
for k in np.arange(start = 1, stop = nTime):
print(" Time[", k, "]=", Time[k])
#
################################################
Time[ 0 ]= 0.0 Time[ 1 ]= 0.125 Time[ 2 ]= 0.25 Time[ 3 ]= 0.375 Time[ 4 ]= 0.5 Time[ 5 ]= 0.625 Time[ 6 ]= 0.75 Time[ 7 ]= 0.875 Time[ 8 ]= 1.0 Time[ 9 ]= 1.125 Time[ 10 ]= 1.25 Time[ 11 ]= 1.375 Time[ 12 ]= 1.5 Time[ 13 ]= 1.625 Time[ 14 ]= 1.75 Time[ 15 ]= 1.875 Time[ 16 ]= 2.0 Time[ 17 ]= 2.125 Time[ 18 ]= 2.25 Time[ 19 ]= 2.375 Time[ 20 ]= 2.5 Time[ 21 ]= 2.625 Time[ 22 ]= 2.75 Time[ 23 ]= 2.875 Time[ 24 ]= 3.0 Time[ 25 ]= 3.125 Time[ 26 ]= 3.25 Time[ 27 ]= 3.375 Time[ 28 ]= 3.5 Time[ 29 ]= 3.625 Time[ 30 ]= 3.75 Time[ 31 ]= 3.875 Time[ 32 ]= 4.0 Time[ 33 ]= 4.125 Time[ 34 ]= 4.25 Time[ 35 ]= 4.375 Time[ 36 ]= 4.5 Time[ 37 ]= 4.625 Time[ 38 ]= 4.75 Time[ 39 ]= 4.875 Time[ 40 ]= 5.0 Time[ 41 ]= 5.125 Time[ 42 ]= 5.25 Time[ 43 ]= 5.375 Time[ 44 ]= 5.5 Time[ 45 ]= 5.625 Time[ 46 ]= 5.75 Time[ 47 ]= 5.875 Time[ 48 ]= 6.0 Time[ 49 ]= 6.125 Time[ 50 ]= 6.25 Time[ 51 ]= 6.375 Time[ 52 ]= 6.5 Time[ 53 ]= 6.625 Time[ 54 ]= 6.75 Time[ 55 ]= 6.875 Time[ 56 ]= 7.0 Time[ 57 ]= 7.125 Time[ 58 ]= 7.25 Time[ 59 ]= 7.375 Time[ 60 ]= 7.5 Time[ 61 ]= 7.625 Time[ 62 ]= 7.75 Time[ 63 ]= 7.875 Time[ 64 ]= 8.0 Time[ 65 ]= 8.125 Time[ 66 ]= 8.25 Time[ 67 ]= 8.375 Time[ 68 ]= 8.5 Time[ 69 ]= 8.625 Time[ 70 ]= 8.75 Time[ 71 ]= 8.875 Time[ 72 ]= 9.0 Time[ 73 ]= 9.125 Time[ 74 ]= 9.25 Time[ 75 ]= 9.375 Time[ 76 ]= 9.5 Time[ 77 ]= 9.625 Time[ 78 ]= 9.75 Time[ 79 ]= 9.875 Time[ 80 ]= 10.0 Time[ 81 ]= 10.125 Time[ 82 ]= 10.25 Time[ 83 ]= 10.375 Time[ 84 ]= 10.5 Time[ 85 ]= 10.625 Time[ 86 ]= 10.75 Time[ 87 ]= 10.875 Time[ 88 ]= 11.0 Time[ 89 ]= 11.125 Time[ 90 ]= 11.25 Time[ 91 ]= 11.375 Time[ 92 ]= 11.5 Time[ 93 ]= 11.625 Time[ 94 ]= 11.75 Time[ 95 ]= 11.875 Time[ 96 ]= 12.0 Time[ 97 ]= 12.125 Time[ 98 ]= 12.25 Time[ 99 ]= 12.375 Time[ 100 ]= 12.5 Time[ 101 ]= 12.625 Time[ 102 ]= 12.75 Time[ 103 ]= 12.875 Time[ 104 ]= 13.0 Time[ 105 ]= 13.125 Time[ 106 ]= 13.25 Time[ 107 ]= 13.375 Time[ 108 ]= 13.5 Time[ 109 ]= 13.625 Time[ 110 ]= 13.75 Time[ 111 ]= 13.875 Time[ 112 ]= 14.0 Time[ 113 ]= 14.125 Time[ 114 ]= 14.25 Time[ 115 ]= 14.375 Time[ 116 ]= 14.5 Time[ 117 ]= 14.625 Time[ 118 ]= 14.75 Time[ 119 ]= 14.875 Time[ 120 ]= 15.0 Time[ 121 ]= 15.125 Time[ 122 ]= 15.25 Time[ 123 ]= 15.375 Time[ 124 ]= 15.5 Time[ 125 ]= 15.625 Time[ 126 ]= 15.75 Time[ 127 ]= 15.875 Time[ 128 ]= 16.0 Time[ 129 ]= 16.125 Time[ 130 ]= 16.25 Time[ 131 ]= 16.375 Time[ 132 ]= 16.5 Time[ 133 ]= 16.625 Time[ 134 ]= 16.75 Time[ 135 ]= 16.875 Time[ 136 ]= 17.0 Time[ 137 ]= 17.125 Time[ 138 ]= 17.25 Time[ 139 ]= 17.375 Time[ 140 ]= 17.5 Time[ 141 ]= 17.625 Time[ 142 ]= 17.75 Time[ 143 ]= 17.875 Time[ 144 ]= 18.0 Time[ 145 ]= 18.125 Time[ 146 ]= 18.25 Time[ 147 ]= 18.375 Time[ 148 ]= 18.5 Time[ 149 ]= 18.625 Time[ 150 ]= 18.75 Time[ 151 ]= 18.875 Time[ 152 ]= 19.0 Time[ 153 ]= 19.125 Time[ 154 ]= 19.25 Time[ 155 ]= 19.375 Time[ 156 ]= 19.5 Time[ 157 ]= 19.625 Time[ 158 ]= 19.75 Time[ 159 ]= 19.875 Time[ 160 ]= 20.0 Time[ 161 ]= 20.125 Time[ 162 ]= 20.25 Time[ 163 ]= 20.375 Time[ 164 ]= 20.5 Time[ 165 ]= 20.625 Time[ 166 ]= 20.75 Time[ 167 ]= 20.875 Time[ 168 ]= 21.0 Time[ 169 ]= 21.125 Time[ 170 ]= 21.25 Time[ 171 ]= 21.375 Time[ 172 ]= 21.5 Time[ 173 ]= 21.625 Time[ 174 ]= 21.75 Time[ 175 ]= 21.875 Time[ 176 ]= 22.0 Time[ 177 ]= 22.125 Time[ 178 ]= 22.25 Time[ 179 ]= 22.375 Time[ 180 ]= 22.5 Time[ 181 ]= 22.625 Time[ 182 ]= 22.75 Time[ 183 ]= 22.875 Time[ 184 ]= 23.0 Time[ 185 ]= 23.125 Time[ 186 ]= 23.25 Time[ 187 ]= 23.375 Time[ 188 ]= 23.5 Time[ 189 ]= 23.625 Time[ 190 ]= 23.75 Time[ 191 ]= 23.875 Time[ 192 ]= 24.0 Time[ 193 ]= 24.125 Time[ 194 ]= 24.25 Time[ 195 ]= 24.375 Time[ 196 ]= 24.5 Time[ 197 ]= 24.625 Time[ 198 ]= 24.75 Time[ 199 ]= 24.875 Time[ 200 ]= 25.0 Time[ 201 ]= 25.125 Time[ 202 ]= 25.25 Time[ 203 ]= 25.375 Time[ 204 ]= 25.5 Time[ 205 ]= 25.625 Time[ 206 ]= 25.75 Time[ 207 ]= 25.875 Time[ 208 ]= 26.0 Time[ 209 ]= 26.125 Time[ 210 ]= 26.25 Time[ 211 ]= 26.375 Time[ 212 ]= 26.5 Time[ 213 ]= 26.625 Time[ 214 ]= 26.75 Time[ 215 ]= 26.875 Time[ 216 ]= 27.0 Time[ 217 ]= 27.125 Time[ 218 ]= 27.25 Time[ 219 ]= 27.375 Time[ 220 ]= 27.5 Time[ 221 ]= 27.625 Time[ 222 ]= 27.75 Time[ 223 ]= 27.875 Time[ 224 ]= 28.0 Time[ 225 ]= 28.125 Time[ 226 ]= 28.25 Time[ 227 ]= 28.375 Time[ 228 ]= 28.5 Time[ 229 ]= 28.625 Time[ 230 ]= 28.75 Time[ 231 ]= 28.875 Time[ 232 ]= 29.0 Time[ 233 ]= 29.125 Time[ 234 ]= 29.25 Time[ 235 ]= 29.375 Time[ 236 ]= 29.5 Time[ 237 ]= 29.625 Time[ 238 ]= 29.75 Time[ 239 ]= 29.875 Time[ 240 ]= 30.0
Victory Lap #6. Insert the equations...¶
Now let's our equations into our loop.
################################################
#
# Step 6: Plumbing Test!
#
k = 0
print(" Time[", k, "]=", Time[k],
" S[", k, "]=", S[k],
" I[", k, "]=", I[k],
" R[", k, "]=", R[k],
" F[", k, "]=", F[k])
for k in np.arange(start = 1, stop = nTime):
S[k] = S[k-1] + dSdt(S[k-1], I[k-1], R[k-1]) * DeltaT
I[k] = I[k-1] + dIdt(S[k-1], I[k-1], R[k-1]) * DeltaT
R[k] = R[k-1] + dRdt(I[k-1]) * DeltaT
F[k] = F[k-1] + dFdt(I[k-1]) * DeltaT
# Because you can have negative personalities
S[k]
print(" Time[", k, "]=", Time[k],
" S[", k, "]=", S[k],
" I[", k, "]=", I[k],
" R[", k, "]=", R[k],
" F[", k, "]=", F[k])
#
################################################
Time[ 0 ]= 0.0 S[ 0 ]= 998.0 I[ 0 ]= 1.0 R[ 0 ]= 0.0 F[ 0 ]= 0.0 Time[ 1 ]= 0.125 S[ 1 ]= 997.8126876876877 I[ 1 ]= 1.1123123123123124 R[ 1 ]= 0.0625 F[ 1 ]= 0.0125 Time[ 2 ]= 0.25 S[ 2 ]= 997.6043743946228 I[ 2 ]= 1.2372021819537025 R[ 2 ]= 0.1320195195195195 F[ 2 ]= 0.026403903903903906 Time[ 3 ]= 0.375 S[ 3 ]= 997.3727169376182 I[ 3 ]= 1.3760694753118352 R[ 3 ]= 0.20934465589162593 F[ 3 ]= 0.041868931178325186 Time[ 4 ]= 0.5 S[ 4 ]= 997.1151133951245 I[ 4 ]= 1.530467807157187 R[ 4 ]= 0.29534899809861564 F[ 4 ]= 0.059069799619723126 Time[ 5 ]= 0.625 S[ 5 ]= 996.8286751781067 I[ 5 ]= 1.7021209386382097 R[ 5 ]= 0.39100323604593984 F[ 5 ]= 0.07820064720918797 Time[ 6 ]= 0.75 S[ 6 ]= 996.5101962388603 I[ 6 ]= 1.8929408074866947 R[ 6 ]= 0.49738579471082794 F[ 6 ]= 0.09947715894216559 Time[ 7 ]= 0.875 S[ 7 ]= 996.1561191612878 I[ 7 ]= 2.105047324497697 R[ 7 ]= 0.6156945951787464 F[ 7 ]= 0.12313891903574928 Time[ 8 ]= 1.0 S[ 8 ]= 995.762497862371 I[ 8 ]= 2.340790074077259 R[ 8 ]= 0.7472600529598524 F[ 8 ]= 0.1494520105919705 Time[ 9 ]= 1.125 S[ 9 ]= 995.3249566226256 I[ 9 ]= 2.6027720582669476 R[ 9 ]= 0.8935594325896812 F[ 9 ]= 0.17871188651793624 Time[ 10 ]= 1.25 S[ 10 ]= 994.8386451541434 I[ 10 ]= 2.8938756223790887 R[ 10 ]= 1.0562326862313653 F[ 10 ]= 0.21124653724627307 Time[ 11 ]= 1.375 S[ 11 ]= 994.2981894096044 I[ 11 ]= 3.217290695239555 R[ 11 ]= 1.2370999126300584 F[ 11 ]= 0.2474199825260117 Time[ 12 ]= 1.5 S[ 12 ]= 993.6976378358681 I[ 12 ]= 3.576545466832903 R[ 12 ]= 1.4381805810825306 F[ 12 ]= 0.28763611621650614 Time[ 13 ]= 1.625 S[ 13 ]= 993.0304027832906 I[ 13 ]= 3.9755396093979716 R[ 13 ]= 1.661714672759587 F[ 13 ]= 0.33234293455191743 Time[ 14 ]= 1.75 S[ 14 ]= 992.2891967990591 I[ 14 ]= 4.418580122924523 R[ 14 ]= 1.9101858983469602 F[ 14 ]= 0.3820371796693921 Time[ 15 ]= 1.875 S[ 15 ]= 991.4659635623993 I[ 15 ]= 4.910419850364987 R[ 15 ]= 2.186347156029743 F[ 15 ]= 0.43726943120594863 Time[ 16 ]= 2.0 S[ 16 ]= 990.5518032649164 I[ 16 ]= 5.456298659070543 R[ 16 ]= 2.4932483966775547 F[ 16 ]= 0.49864967933551096 Time[ 17 ]= 2.125 S[ 17 ]= 989.5368923047012 I[ 17 ]= 6.061987219855509 R[ 17 ]= 2.8342670628694635 F[ 17 ]= 0.5668534125738928 Time[ 18 ]= 2.25 S[ 18 ]= 988.410397253074 I[ 18 ]= 6.73383322999348 R[ 18 ]= 3.213141264110433 F[ 18 ]= 0.6426282528220866 Time[ 19 ]= 2.375 S[ 19 ]= 987.1603831737759 I[ 19 ]= 7.478809817042138 R[ 19 ]= 3.6340058409850253 F[ 19 ]= 0.726801168197005 Time[ 20 ]= 2.5 S[ 20 ]= 985.7737165328239 I[ 20 ]= 8.304565721716058 R[ 20 ]= 4.101431454550159 F[ 20 ]= 0.8202862909100317 Time[ 21 ]= 2.625 S[ 21 ]= 984.2359631409645 I[ 21 ]= 9.219476684446692 R[ 21 ]= 4.620466812157413 F[ 21 ]= 0.9240933624314824 Time[ 22 ]= 2.75 S[ 22 ]= 982.5312818285589 I[ 22 ]= 10.232697245518711 R[ 22 ]= 5.196684104935331 F[ 22 ]= 1.0393368209870661 Time[ 23 ]= 2.875 S[ 23 ]= 980.6423148747208 I[ 23 ]= 11.354211905942972 R[ 23 ]= 5.83622768278025 F[ 23 ]= 1.1672455365560501 Time[ 24 ]= 3.0 S[ 24 ]= 978.5500766093621 I[ 24 ]= 12.59488427835589 R[ 24 ]= 6.545865926901686 F[ 24 ]= 1.3091731853803372 Time[ 25 ]= 3.125 S[ 25 ]= 976.2338420898177 I[ 25 ]= 13.966502477023633 R[ 25 ]= 7.333046194298929 F[ 25 ]= 1.4666092388597858 Time[ 26 ]= 3.25 S[ 26 ]= 973.6710383343279 I[ 26 ]= 15.481818546736612 R[ 26 ]= 8.205952599112907 F[ 26 ]= 1.6411905198225811 Time[ 27 ]= 3.375 S[ 27 ]= 970.8371412836088 I[ 27 ]= 17.15457920645051 R[ 27 ]= 9.173566258283945 F[ 27 ]= 1.8347132516567888 Time[ 28 ]= 3.5 S[ 28 ]= 967.7055824679519 I[ 28 ]= 18.999544581623617 R[ 28 ]= 10.245727458687101 F[ 28 ]= 2.0491454917374203 Time[ 29 ]= 3.625 S[ 29 ]= 964.2476702863652 I[ 29 ]= 21.03249091958851 R[ 29 ]= 11.433198995038577 F[ 29 ]= 2.2866397990077156 Time[ 30 ]= 3.75 S[ 30 ]= 960.4325318562747 I[ 30 ]= 23.27019253070975 R[ 30 ]= 12.747729677512858 F[ 30 ]= 2.549545935502572 Time[ 31 ]= 3.875 S[ 31 ]= 956.2270825591177 I[ 31 ]= 25.730377388063555 R[ 31 ]= 14.202116710682217 F[ 31 ]= 2.840423342136444 Time[ 32 ]= 4.0 S[ 32 ]= 951.5960316688802 I[ 32 ]= 28.431649974196237 R[ 32 ]= 15.810265297436189 F[ 32 ]= 3.1620530594872385 Time[ 33 ]= 4.125 S[ 33 ]= 946.5019337714075 I[ 33 ]= 31.39337412360423 R[ 33 ]= 17.587243420823455 F[ 33 ]= 3.5174486841646915 Time[ 34 ]= 4.25 S[ 34 ]= 940.9052970053377 I[ 34 ]= 34.63550783040367 R[ 34 ]= 19.54932930354872 F[ 34 ]= 3.9098658607097443 Time[ 35 ]= 4.375 S[ 35 ]= 934.764760397638 I[ 35 ]= 38.1783813508231 R[ 35 ]= 21.71404854294895 F[ 35 ]= 4.34280970858979 Time[ 36 ]= 4.5 S[ 36 ]= 928.0373536128977 I[ 36 ]= 42.04240953425173 R[ 36 ]= 24.100197377375395 F[ 36 ]= 4.820039475475079 Time[ 37 ]= 4.625 S[ 37 ]= 920.6788531339405 I[ 37 ]= 46.24772929814006 R[ 37 ]= 26.72784797326613 F[ 37 ]= 5.345569594653226 Time[ 38 ]= 4.75 S[ 38 ]= 912.6442490501374 I[ 38 ]= 50.813753684582665 R[ 38 ]= 29.61833105439988 F[ 38 ]= 5.9236662108799765 Time[ 39 ]= 4.875 S[ 39 ]= 903.8883360172605 I[ 39 ]= 55.758635191115964 R[ 39 ]= 32.7941906596863 F[ 39 ]= 6.55883813193726 Time[ 40 ]= 5.0 S[ 40 ]= 894.3664403023124 I[ 40 ]= 61.09863326673036 R[ 40 ]= 36.27910535913105 F[ 40 ]= 7.25582107182621 Time[ 41 ]= 5.125 S[ 41 ]= 884.0352918500802 I[ 41 ]= 66.84738422395782 R[ 41 ]= 40.097769938301695 F[ 41 ]= 8.01955398766034 Time[ 42 ]= 5.25 S[ 42 ]= 872.854045717846 I[ 42 ]= 73.01507653939517 R[ 42 ]= 44.275731452299055 F[ 42 ]= 8.855146290459812 Time[ 43 ]= 5.375 S[ 43 ]= 860.7854507698995 I[ 43 ]= 79.60754074688708 R[ 43 ]= 48.839173736011254 F[ 43 ]= 9.76783474720225 Time[ 44 ]= 5.5 S[ 44 ]= 847.7971550391052 I[ 44 ]= 86.62527092166485 R[ 44 ]= 53.8146450326917 F[ 44 ]= 10.76292900653834 Time[ 45 ]= 5.625 S[ 45 ]= 833.8631266318229 I[ 45 ]= 94.06240400982227 R[ 45 ]= 59.228724465295755 F[ 45 ]= 11.84574489305915 Time[ 46 ]= 5.75 S[ 46 ]= 818.9651566779708 I[ 46 ]= 101.90569366293776 R[ 46 ]= 65.10762471590965 F[ 46 ]= 13.021524943181928 Time[ 47 ]= 5.875 S[ 47 ]= 803.0943971060701 I[ 47 ]= 110.13352621011803 R[ 47 ]= 71.47673056984326 F[ 47 ]= 14.29534611396865 Time[ 48 ]= 6.0 S[ 48 ]= 786.25287180382 I[ 48 ]= 118.71503704660932 R[ 48 ]= 78.36007595797564 F[ 48 ]= 15.672015191595126 Time[ 49 ]= 6.125 S[ 49 ]= 768.4548862423735 I[ 49 ]= 127.60939482956013 R[ 49 ]= 85.77976577338872 F[ 49 ]= 17.155953154677743 Time[ 50 ]= 6.25 S[ 50 ]= 749.7282494927003 I[ 50 ]= 136.7653269670163 R[ 50 ]= 93.75535295023623 F[ 50 ]= 18.751070590047245 Time[ 51 ]= 6.375 S[ 51 ]= 730.1152156049717 I[ 51 ]= 146.1209613322187 R[ 51 ]= 102.30318588567475 F[ 51 ]= 20.46063717713495 Time[ 52 ]= 6.5 S[ 52 ]= 709.6730504937525 I[ 52 ]= 155.6040543435215 R[ 52 ]= 111.43574596893842 F[ 52 ]= 22.287149193787684 Time[ 53 ]= 6.625 S[ 53 ]= 688.4741375102669 I[ 53 ]= 165.13266325124292 R[ 53 ]= 121.16099936540851 F[ 53 ]= 24.232199873081704 Time[ 54 ]= 6.75 S[ 54 ]= 666.6055509815309 I[ 54 ]= 174.6163000361357 R[ 54 ]= 131.4817908186112 F[ 54 ]= 26.29635816372224 Time[ 55 ]= 6.875 S[ 55 ]= 644.1680524421113 I[ 55 ]= 183.95757607284511 R[ 55 ]= 142.39530957086967 F[ 55 ]= 28.479061914173936 Time[ 56 ]= 7.0 S[ 56 ]= 621.2744981545549 I[ 56 ]= 193.05431215493815 R[ 56 ]= 153.8926580754225 F[ 56 ]= 30.7785316150845 Time[ 57 ]= 7.125 S[ 57 ]= 598.0476865177692 I[ 57 ]= 201.80205038010348 R[ 57 ]= 165.95855258510613 F[ 57 ]= 33.19171051702123 Time[ 58 ]= 7.25 S[ 58 ]= 574.6177165100238 I[ 58 ]= 210.09686660934113 R[ 58 ]= 178.5711807338626 F[ 58 ]= 35.71423614677252 Time[ 59 ]= 7.375 S[ 59 ]= 551.1189688641166 I[ 59 ]= 217.8383492595478 R[ 59 ]= 191.70223489694644 F[ 59 ]= 38.340446979389284 Time[ 60 ]= 7.5 S[ 60 ]= 527.6868553401007 I[ 60 ]= 224.93258658909767 R[ 60 ]= 205.31713172566816 F[ 60 ]= 41.063426345133635 Time[ 61 ]= 7.625 S[ 61 ]= 504.4545037917899 I[ 61 ]= 231.29499414322618 R[ 61 ]= 219.37541838748677 F[ 61 ]= 43.87508367749736 Time[ 62 ]= 7.75 S[ 62 ]= 481.5495544941718 I[ 62 ]= 236.8528188801023 R[ 62 ]= 233.8313555214384 F[ 62 ]= 46.76627110428768 Time[ 63 ]= 7.875 S[ 63 ]= 459.09123505444154 I[ 63 ]= 241.54717690382483 R[ 63 ]= 248.6346567014448 F[ 63 ]= 49.726931340288964 Time[ 64 ]= 8.0 S[ 64 ]= 437.18785799352236 I[ 64 ]= 245.33451569695717 R[ 64 ]= 263.73135525793384 F[ 64 ]= 52.746271051586774 Time[ 65 ]= 8.125 S[ 65 ]= 415.93484965756824 I[ 65 ]= 248.18743535563948 R[ 65 ]= 279.06476248899367 F[ 65 ]= 55.81295249779874 Time[ 66 ]= 8.25 S[ 66 ]= 395.4133759706434 I[ 66 ]= 250.09485139089136 R[ 66 ]= 294.57647719872114 F[ 66 ]= 58.915295439744234 Time[ 67 ]= 8.375 S[ 67 ]= 375.6895849212227 I[ 67 ]= 251.06152858599523 R[ 67 ]= 310.20740541065186 F[ 67 ]= 62.041481082130375 Time[ 68 ]= 8.5 S[ 68 ]= 356.814442727142 I[ 68 ]= 251.1070561361263 R[ 68 ]= 325.89875094727654 F[ 68 ]= 65.17975018945532 Time[ 69 ]= 8.625 S[ 69 ]= 338.82410458693727 I[ 69 ]= 250.2643650661216 R[ 69 ]= 341.59294195578445 F[ 69 ]= 68.3185883911569 Time[ 70 ]= 8.75 S[ 70 ]= 321.74073460678414 I[ 70 ]= 248.57790766631564 R[ 70 ]= 357.234464772417 F[ 70 ]= 71.44689295448342 Time[ 71 ]= 8.875 S[ 71 ]= 305.57367406085365 I[ 71 ]= 246.10162513727246 R[ 71 ]= 372.77058400156176 F[ 71 ]= 74.55411680031236 Time[ 72 ]= 9.0 S[ 72 ]= 290.3208522450594 I[ 72 ]= 242.89682506777126 R[ 72 ]= 388.15193557264126 F[ 72 ]= 77.63038711452826 Time[ 73 ]= 9.125 S[ 73 ]= 275.97033829107227 I[ 73 ]= 239.03007714167558 R[ 73 ]= 403.332987139377 F[ 73 ]= 80.6665974278754 Time[ 74 ]= 9.25 S[ 74 ]= 262.50194318006805 I[ 74 ]= 234.5712164670541 R[ 74 ]= 418.27236696073174 F[ 74 ]= 83.65447339214634 Time[ 75 ]= 9.375 S[ 75 ]= 249.88879635187573 I[ 75 ]= 229.59152206021736 R[ 75 ]= 432.9330679899226 F[ 75 ]= 86.58661359798452 Time[ 76 ]= 9.5 S[ 76 ]= 238.0988384045117 I[ 76 ]= 224.1621158530651 R[ 76 ]= 447.2825381186862 F[ 76 ]= 89.45650762373724 Time[ 77 ]= 9.625 S[ 77 ]= 227.09618848048126 I[ 77 ]= 218.35260708811566 R[ 77 ]= 461.2926703595028 F[ 77 ]= 92.25853407190056 Time[ 78 ]= 9.75 S[ 78 ]= 216.8423606231661 I[ 78 ]= 212.22998941382215 R[ 78 ]= 474.93970830251 F[ 78 ]= 94.987941660502 Time[ 79 ]= 9.875 S[ 79 ]= 207.29731678048762 I[ 79 ]= 205.85778405046395 R[ 79 ]= 488.2040826408739 F[ 79 ]= 97.64081652817478 Time[ 80 ]= 10.0 S[ 80 ]= 198.42035482932857 I[ 80 ]= 199.29541219783817 R[ 80 ]= 501.0701941440279 F[ 80 ]= 100.21403882880558 Time[ 81 ]= 10.125 S[ 81 ]= 190.17083795757327 I[ 81 ]= 192.5977731547556 R[ 81 ]= 513.5261574063927 F[ 81 ]= 102.70523148127856 Time[ 82 ]= 10.25 S[ 82 ]= 182.50877718888205 I[ 82 ]= 185.81500093684014 R[ 82 ]= 525.5635182285649 F[ 82 ]= 105.112703645713 Time[ 83 ]= 10.375 S[ 83 ]= 175.395282132798 I[ 83 ]= 178.99237092266117 R[ 83 ]= 537.1769557871174 F[ 83 ]= 107.4353911574235 Time[ 84 ]= 10.5 S[ 84 ]= 168.79289661449945 I[ 84 ]= 172.17032862176012 R[ 84 ]= 548.3639789697837 F[ 84 ]= 109.67279579395677 Time[ 85 ]= 10.625 S[ 85 ]= 162.66583610845902 I[ 85 ]= 165.38461448116857 R[ 85 ]= 559.1246245086437 F[ 85 ]= 111.82492490172876 Time[ 86 ]= 10.75 S[ 86 ]= 156.98014325259686 I[ 86 ]= 158.6664612509431 R[ 86 ]= 569.4611629137167 F[ 86 ]= 113.89223258274338 Time[ 87 ]= 10.875 S[ 87 ]= 151.70377647837716 I[ 87 ]= 152.04284343134205 R[ 87 ]= 579.3778167419007 F[ 87 ]= 115.87556334838017 Time[ 88 ]= 11.0 S[ 88 ]= 146.8066452154054 I[ 88 ]= 145.53676143696316 R[ 88 ]= 588.8804944563595 F[ 88 ]= 117.77609889127194 Time[ 89 ]= 11.125 S[ 89 ]= 142.26060341025374 I[ 89 ]= 139.16754613434262 R[ 89 ]= 597.9765420461697 F[ 89 ]= 119.59530840923398 Time[ 90 ]= 11.25 S[ 90 ]= 138.03941137599043 I[ 90 ]= 132.95117220853024 R[ 90 ]= 606.6745136795661 F[ 90 ]= 121.33490273591326 Time[ 91 ]= 11.375 S[ 91 ]= 134.11867435227623 I[ 91 ]= 126.90057131660467 R[ 91 ]= 614.9839619425992 F[ 91 ]= 122.99679238851989 Time[ 92 ]= 11.5 S[ 92 ]= 130.47576466064058 I[ 92 ]= 121.02593815949497 R[ 92 ]= 622.915247649887 F[ 92 ]= 124.58304952997744 Time[ 93 ]= 11.625 S[ 93 ]= 127.08973301335762 I[ 93 ]= 115.33502444481582 R[ 93 ]= 630.4793687848554 F[ 93 ]= 126.09587375697113 Time[ 94 ]= 11.75 S[ 94 ]= 123.94121338579235 I[ 94 ]= 109.8334172390199 R[ 94 ]= 637.6878078126564 F[ 94 ]= 127.53756156253132 Time[ 95 ]= 11.875 S[ 95 ]= 121.01232488702641 I[ 95 ]= 104.52479944485934 R[ 95 ]= 644.5523963900952 F[ 95 ]= 128.91047927801907 Time[ 96 ]= 12.0 S[ 96 ]= 118.28657324999725 I[ 96 ]= 99.41119112352406 R[ 96 ]= 651.0851963553989 F[ 96 ]= 130.2170392710798 Time[ 97 ]= 12.125 S[ 97 ]= 115.74875389401932 I[ 97 ]= 94.49317114523768 R[ 97 ]= 657.2983958006191 F[ 97 ]= 131.45967916012387 Time[ 98 ]= 12.25 S[ 98 ]= 113.38485797137068 I[ 98 ]= 89.77007923199349 R[ 98 ]= 663.2042189971964 F[ 98 ]= 132.64084379943935 Time[ 99 ]= 12.375 S[ 99 ]= 111.18198237750974 I[ 99 ]= 85.24019888345492 R[ 99 ]= 668.8148489491961 F[ 99 ]= 133.76296978983927 Time[ 100 ]= 12.5 S[ 100 ]= 109.12824436427991 I[ 100 ]= 80.90092198042564 R[ 100 ]= 674.142361379412 F[ 100 ]= 134.82847227588246 Time[ 101 ]= 12.625 S[ 101 ]= 107.2127011315551 I[ 101 ]= 76.74889606461852 R[ 101 ]= 679.1986690031886 F[ 101 ]= 135.8397338006378 Time[ 102 ]= 12.75 S[ 102 ]= 105.42527457138164 I[ 102 ]= 72.7801554199456 R[ 102 ]= 683.9954750072272 F[ 102 ]= 136.79909500144552 Time[ 103 ]= 12.875 S[ 103 ]= 103.75668118786467 I[ 103 ]= 68.99023714696665 R[ 103 ]= 688.5442347209738 F[ 103 ]= 137.70884694419485 Time[ 104 ]= 13.0 S[ 104 ]= 102.1983671057154 I[ 104 ]= 65.37428344309342 R[ 104 ]= 692.8561245426592 F[ 104 ]= 138.57122490853192 Time[ 105 ]= 13.125 S[ 105 ]= 100.74244800206557 I[ 105 ]= 61.92713128851124 R[ 105 ]= 696.9420172578526 F[ 105 ]= 139.38840345157058 Time[ 106 ]= 13.25 S[ 106 ]= 99.38165374289086 I[ 106 ]= 58.64339070104761 R[ 106 ]= 700.8124629633846 F[ 106 ]= 140.16249259267698 Time[ 107 ]= 13.375 S[ 107 ]= 98.1092774714714 I[ 107 ]= 55.51751266988849 R[ 107 ]= 704.4776748822001 F[ 107 ]= 140.89553497644008 Time[ 108 ]= 13.5 S[ 108 ]= 96.91912887715665 I[ 108 ]= 52.543847813961605 R[ 108 ]= 707.9475194240681 F[ 108 ]= 141.5895038848137 Time[ 109 ]= 13.625 S[ 109 ]= 95.80549136461383 I[ 109 ]= 49.71669674045731 R[ 109 ]= 711.2315099124407 F[ 109 ]= 142.2463019824882 Time[ 110 ]= 13.75 S[ 110 ]= 94.76308284380733 I[ 110 ]= 47.030353005729516 R[ 110 ]= 714.3388034587193 F[ 110 ]= 142.8677606917439 Time[ 111 ]= 13.875 S[ 111 ]= 93.7870198668885 I[ 111 ]= 44.479139507218626 R[ 111 ]= 717.2782005215774 F[ 111 ]= 143.45564010431553 Time[ 112 ]= 14.0 S[ 112 ]= 92.87278484818921 I[ 112 ]= 42.05743906287652 R[ 112 ]= 720.0581467407786 F[ 112 ]= 144.01162934815576 Time[ 113 ]= 14.125 S[ 113 ]= 92.01619611623347 I[ 113 ]= 39.75971986511652 R[ 113 ]= 722.6867366822084 F[ 113 ]= 144.5373473364417 Time[ 114 ]= 14.25 S[ 114 ]= 91.2133805610647 I[ 114 ]= 37.58055643040156 R[ 114 ]= 725.1717191737782 F[ 114 ]= 145.03434383475567 Time[ 115 ]= 14.375 S[ 115 ]= 90.46074865544401 I[ 115 ]= 35.51464660374213 R[ 115 ]= 727.5205039506783 F[ 115 ]= 145.50410079013568 Time[ 116 ]= 14.5 S[ 116 ]= 89.75497164402641 I[ 116 ]= 33.55682511987906 R[ 116 ]= 729.7401693634122 F[ 116 ]= 145.94803387268246 Time[ 117 ]= 14.625 S[ 117 ]= 89.09296071004464 I[ 117 ]= 31.7020741698699 R[ 117 ]= 731.8374709334046 F[ 117 ]= 146.36749418668094 Time[ 118 ]= 14.75 S[ 118 ]= 88.47184794402602 I[ 118 ]= 29.945531373148267 R[ 118 ]= 733.8188505690215 F[ 118 ]= 146.76377011380433 Time[ 119 ]= 14.875 S[ 119 ]= 87.88896895343456 I[ 119 ]= 28.282495510753616 R[ 119 ]= 735.6904462798433 F[ 119 ]= 147.13808925596868 Time[ 120 ]= 15.0 S[ 120 ]= 87.34184696573634 I[ 120 ]= 26.708430335145312 R[ 120 ]= 737.4581022492654 F[ 120 ]= 147.4916204498531 Time[ 121 ]= 15.125 S[ 121 ]= 86.82817829015772 I[ 121 ]= 25.21896673558803 R[ 121 ]= 739.127379145212 F[ 121 ]= 147.8254758290424 Time[ 122 ]= 15.25 S[ 122 ]= 86.34581901530514 I[ 122 ]= 23.809903505271514 R[ 122 ]= 740.7035645661863 F[ 122 ]= 148.14071291323725 Time[ 123 ]= 15.375 S[ 123 ]= 85.89277283083912 I[ 123 ]= 22.477206926842168 R[ 123 ]= 742.1916835352657 F[ 123 ]= 148.43833670705314 Time[ 124 ]= 15.5 S[ 124 ]= 85.46717987155753 I[ 124 ]= 21.217009366610593 R[ 124 ]= 743.5965089681933 F[ 124 ]= 148.71930179363866 Time[ 125 ]= 15.625 S[ 125 ]= 85.0673064915745 I[ 125 ]= 20.02560704409784 R[ 125 ]= 744.9225720536065 F[ 125 ]= 148.9845144107213 Time[ 126 ]= 15.75 S[ 126 ]= 84.69153588482216 I[ 126 ]= 18.89945712254284 R[ 126 ]= 746.1741724938626 F[ 126 ]= 149.2348344987725 Time[ 127 ]= 15.875 S[ 127 ]= 84.33835947589775 I[ 127 ]= 17.835174247276537 R[ 127 ]= 747.3553885640215 F[ 127 ]= 149.4710777128043 Time[ 128 ]= 16.0 S[ 128 ]= 84.00636901237732 I[ 128 ]= 16.82952664225123 R[ 128 ]= 748.4700869544763 F[ 128 ]= 149.69401739089525 Time[ 129 ]= 16.125 S[ 129 ]= 83.69424929617033 I[ 129 ]= 15.879431860289388 R[ 129 ]= 749.521932369617 F[ 129 ]= 149.90438647392338 Time[ 130 ]= 16.25 S[ 130 ]= 83.4007714973456 I[ 130 ]= 14.981952269592412 R[ 130 ]= 750.5143968608851 F[ 130 ]= 150.102879372177 Time[ 131 ]= 16.375 S[ 131 ]= 83.12478699916747 I[ 131 ]= 14.13429034755111 R[ 131 ]= 751.4507688777346 F[ 131 ]= 150.2901537755469 Time[ 132 ]= 16.5 S[ 132 ]= 82.86522172788735 I[ 132 ]= 13.333783842764895 R[ 132 ]= 752.3341620244565 F[ 132 ]= 150.4668324048913 Time[ 133 ]= 16.625 S[ 133 ]= 82.62107092518494 I[ 133 ]= 12.577900857259928 R[ 133 ]= 753.1675235146294 F[ 133 ]= 150.63350470292588 Time[ 134 ]= 16.75 S[ 134 ]= 82.39139432508527 I[ 134 ]= 11.864234893065102 R[ 134 ]= 753.9536423182082 F[ 134 ]= 150.7907284636416 Time[ 135 ]= 16.875 S[ 135 ]= 82.17531170073124 I[ 135 ]= 11.190499900439246 R[ 135 ]= 754.6951569990247 F[ 135 ]= 150.93903139980492 Time[ 136 ]= 17.0 S[ 136 ]= 81.9719987496021 I[ 136 ]= 10.55452535903545 R[ 136 ]= 755.3945632428022 F[ 136 ]= 151.07891264856042 Time[ 137 ]= 17.125 S[ 137 ]= 81.78068328866782 I[ 137 ]= 9.954251418042064 R[ 137 ]= 756.0542210777419 F[ 137 ]= 151.21084421554835 Time[ 138 ]= 17.25 S[ 138 ]= 81.60064173358887 I[ 138 ]= 9.387724116767856 R[ 138 ]= 756.6763617913695 F[ 138 ]= 151.33527235827387 Time[ 139 ]= 17.375 S[ 139 ]= 81.43119583843584 I[ 139 ]= 8.853090703163298 R[ 139 ]= 757.2630945486675 F[ 139 ]= 151.45261890973347 Time[ 140 ]= 17.5 S[ 140 ]= 81.27170967454049 I[ 140 ]= 8.3485950643214 R[ 140 ]= 757.8164127176152 F[ 140 ]= 151.563282543523 Time[ 141 ]= 17.625 S[ 141 ]= 81.12158682901972 I[ 141 ]= 7.8725732800180594 R[ 141 ]= 758.3381999091353 F[ 141 ]= 151.66763998182702 Time[ 142 ]= 17.75 S[ 142 ]= 80.98026780525824 I[ 142 ]= 7.423449307778181 R[ 142 ]= 758.8302357391364 F[ 142 ]= 151.76604714782724 Time[ 143 ]= 17.875 S[ 143 ]= 80.84722760921228 I[ 143 ]= 6.999730805740775 R[ 143 ]= 759.2942013208725 F[ 143 ]= 151.85884026417446 Time[ 144 ]= 18.0 S[ 144 ]= 80.72197350682232 I[ 144 ]= 6.600005097700178 R[ 144 ]= 759.7316844962313 F[ 144 ]= 151.94633689924623 Time[ 145 ]= 18.125 S[ 145 ]= 80.60404293911236 I[ 145 ]= 6.222935283082623 R[ 145 ]= 760.1441848148376 F[ 145 ]= 152.0288369629675 Time[ 146 ]= 18.25 S[ 146 ]= 80.49300158272051 I[ 146 ]= 5.867256493243283 R[ 146 ]= 760.5331182700302 F[ 146 ]= 152.10662365400603 Time[ 147 ]= 18.375 S[ 147 ]= 80.38844154466238 I[ 147 ]= 5.531772294308162 R[ 147 ]= 760.8998218008579 F[ 147 ]= 152.17996436017157 Time[ 148 ]= 18.5 S[ 148 ]= 80.28997968108663 I[ 148 ]= 5.215351235810802 R[ 148 ]= 761.2455575692521 F[ 148 ]= 152.2491115138504 Time[ 149 ]= 18.625 S[ 149 ]= 80.19725603064967 I[ 149 ]= 4.916923543561962 R[ 149 ]= 761.5715170214903 F[ 149 ]= 152.31430340429804 Time[ 150 ]= 18.75 S[ 150 ]= 80.10993235392444 I[ 150 ]= 4.635477954520054 R[ 150 ]= 761.8788247429629 F[ 150 ]= 152.37576494859258 Time[ 151 ]= 18.875 S[ 151 ]= 80.02769077097275 I[ 151 ]= 4.37005869088273 R[ 151 ]= 762.1685421151204 F[ 151 ]= 152.4337084230241 Time[ 152 ]= 19.0 S[ 152 ]= 79.95023248986014 I[ 152 ]= 4.119762570179137 R[ 152 ]= 762.4416707833007 F[ 152 ]= 152.48833415666013 Time[ 153 ]= 19.125 S[ 153 ]= 79.87727661948223 I[ 153 ]= 3.8837362477936157 R[ 153 ]= 762.6991559439368 F[ 153 ]= 152.53983118878736 Time[ 154 ]= 19.25 S[ 154 ]= 79.80855906060899 I[ 154 ]= 3.661173588082347 R[ 154 ]= 762.9418894594239 F[ 154 ]= 152.5883778918848 Time[ 155 ]= 19.375 S[ 155 ]= 79.74383146954168 I[ 155 ]= 3.4513131600434748 R[ 155 ]= 763.170712808679 F[ 155 ]= 152.6341425617358 Time[ 156 ]= 19.5 S[ 156 ]= 79.68286028922313 I[ 156 ]= 3.253435853358767 R[ 156 ]= 763.3864198811817 F[ 156 ]= 152.67728397623637 Time[ 157 ]= 19.625 S[ 157 ]= 79.62542584304761 I[ 157 ]= 3.0668626105323806 R[ 157 ]= 763.5897596220167 F[ 157 ]= 152.71795192440337 Time[ 158 ]= 19.75 S[ 158 ]= 79.57132148698783 I[ 158 ]= 2.89095227080223 R[ 158 ]= 763.781438535175 F[ 158 ]= 152.75628770703503 Time[ 159 ]= 19.875 S[ 159 ]= 79.52035281599464 I[ 159 ]= 2.7250995214852507 R[ 159 ]= 763.9621230521001 F[ 159 ]= 152.79242461042006 Time[ 160 ]= 20.0 S[ 160 ]= 79.47233692093468 I[ 160 ]= 2.5687329524338187 R[ 160 ]= 764.1324417721929 F[ 160 ]= 152.82648835443862 Time[ 161 ]= 20.125 S[ 161 ]= 79.4271016926143 I[ 161 ]= 2.421313209321667 R[ 161 ]= 764.29298758172 F[ 161 ]= 152.85859751634405 Time[ 162 ]= 20.25 S[ 162 ]= 79.38448516969731 I[ 162 ]= 2.2823312415395356 R[ 162 ]= 764.4443196573026 F[ 162 ]= 152.88886393146058 Time[ 163 ]= 20.375 S[ 163 ]= 79.34433492756168 I[ 163 ]= 2.151306640559709 R[ 163 ]= 764.5869653598988 F[ 163 ]= 152.91739307197983 Time[ 164 ]= 20.5 S[ 164 ]= 79.30650750535814 I[ 164 ]= 2.027786064721268 R[ 164 ]= 764.7214220249339 F[ 164 ]= 152.94428440498683 Time[ 165 ]= 20.625 S[ 165 ]= 79.27086786873386 I[ 165 ]= 1.911341746491454 R[ 165 ]= 764.848158653979 F[ 165 ]= 152.96963173079584 Time[ 166 ]= 20.75 S[ 166 ]= 79.23728890586783 I[ 166 ]= 1.8015700783706272 R[ 166 ]= 764.9676175131348 F[ 166 ]= 152.99352350262697 Time[ 167 ]= 20.875 S[ 167 ]= 79.20565095463391 I[ 167 ]= 1.6980902737267496 R[ 167 ]= 765.080215643033 F[ 167 ]= 153.0160431286066 Time[ 168 ]= 21.0 S[ 168 ]= 79.17584135886281 I[ 168 ]= 1.6005430989683453 R[ 168 ]= 765.1863462851409 F[ 168 ]= 153.03726925702819 Time[ 169 ]= 21.125 S[ 169 ]= 79.14775405181753 I[ 169 ]= 1.5085896735909965 R[ 169 ]= 765.2863802288264 F[ 169 ]= 153.0572760457653 Time[ 170 ]= 21.25 S[ 170 ]= 79.12128916512896 I[ 170 ]= 1.4219103347602395 R[ 170 ]= 765.3806670834258 F[ 170 ]= 153.07613341668517 Time[ 171 ]= 21.375 S[ 171 ]= 79.09635266155996 I[ 171 ]= 1.3402035632222096 R[ 171 ]= 765.4695364793483 F[ 171 ]= 153.09390729586966 Time[ 172 ]= 21.5 S[ 172 ]= 79.07285599007896 I[ 172 ]= 1.2631849674615427 R[ 172 ]= 765.5532992020497 F[ 172 ]= 153.11065984040994 Time[ 173 ]= 21.625 S[ 173 ]= 79.05071576182773 I[ 173 ]= 1.1905863231531648 R[ 173 ]= 765.6322482625161 F[ 173 ]= 153.1264496525032 Time[ 174 ]= 21.75 S[ 174 ]= 79.02985344566441 I[ 174 ]= 1.1221546650799878 R[ 174 ]= 765.7066599077132 F[ 174 ]= 153.1413319815426 Time[ 175 ]= 21.875 S[ 175 ]= 79.01019508205171 I[ 175 ]= 1.0576514288116994 R[ 175 ]= 765.7767945742806 F[ 175 ]= 153.1553589148561 Time[ 176 ]= 22.0 S[ 176 ]= 78.9916710141422 I[ 176 ]= 0.9968516395603295 R[ 176 ]= 765.8428977885814 F[ 176 ]= 153.16857955771624 Time[ 177 ]= 22.125 S[ 177 ]= 78.97421563498973 I[ 177 ]= 0.9395431457457671 R[ 177 ]= 765.9052010160539 F[ 177 ]= 153.18104020321076 Time[ 178 ]= 22.25 S[ 178 ]= 78.95776714988594 I[ 178 ]= 0.885525894918623 R[ 178 ]= 765.9639224626629 F[ 178 ]= 153.19278449253258 Time[ 179 ]= 22.375 S[ 179 ]= 78.94226735288709 I[ 179 ]= 0.8346112497985825 R[ 179 ]= 766.0192678310954 F[ 179 ]= 153.20385356621907 Time[ 180 ]= 22.5 S[ 180 ]= 78.92766141665726 I[ 180 ]= 0.78662134229352 R[ 180 ]= 766.0714310342078 F[ 180 ]= 153.21428620684154 Time[ 181 ]= 22.625 S[ 181 ]= 78.9138976948107 I[ 181 ]= 0.7413884634680576 R[ 181 ]= 766.1205948681011 F[ 181 ]= 153.2241189736202 Time[ 182 ]= 22.75 S[ 182 ]= 78.90092753598877 I[ 182 ]= 0.6987544875298815 R[ 182 ]= 766.1669316470678 F[ 182 ]= 153.23338632941355 Time[ 183 ]= 22.875 S[ 183 ]= 78.88870510895595 I[ 183 ]= 0.658570327997959 R[ 183 ]= 766.2106038025385 F[ 183 ]= 153.24212076050767 Time[ 184 ]= 23.0 S[ 184 ]= 78.87718723804522 I[ 184 ]= 0.6206954243088277 R[ 184 ]= 766.2517644480383 F[ 184 ]= 153.25035288960765 Time[ 185 ]= 23.125 S[ 185 ]= 78.8663332483255 I[ 185 ]= 0.5849972572053803 R[ 185 ]= 766.2905579120576 F[ 185 ]= 153.25811158241152 Time[ 186 ]= 23.25 S[ 186 ]= 78.85610481990338 I[ 186 ]= 0.5513508913371014 R[ 186 ]= 766.3271202406329 F[ 186 ]= 153.2654240481266 Time[ 187 ]= 23.375 S[ 187 ]= 78.84646585080863 I[ 187 ]= 0.5196385435815755 R[ 187 ]= 766.3615796713415 F[ 187 ]= 153.2723159342683 Time[ 188 ]= 23.5 S[ 188 ]= 78.83738232794722 I[ 188 ]= 0.48974917567436665 R[ 188 ]= 766.3940570803153 F[ 188 ]= 153.27881141606306 Time[ 189 ]= 23.625 S[ 189 ]= 78.82882220563786 I[ 189 ]= 0.46157810980814756 R[ 189 ]= 766.424666403795 F[ 189 ]= 153.284933280759 Time[ 190 ]= 23.75 S[ 190 ]= 78.82075529127806 I[ 190 ]= 0.43502666593233474 R[ 190 ]= 766.4535150356579 F[ 190 ]= 153.29070300713158 Time[ 191 ]= 23.875 S[ 191 ]= 78.81315313771391 I[ 191 ]= 0.4100018195515591 R[ 191 ]= 766.4807042022787 F[ 191 ]= 153.29614084045573 Time[ 192 ]= 24.0 S[ 192 ]= 78.80598894191392 I[ 192 ]= 0.38641587888517903 R[ 192 ]= 766.5063293160007 F[ 192 ]= 153.30126586320014 Time[ 193 ]= 24.125 S[ 193 ]= 78.79923744957188 I[ 193 ]= 0.3641861803108347 R[ 193 ]= 766.530480308431 F[ 193 ]= 153.30609606168622 Time[ 194 ]= 24.25 S[ 194 ]= 78.79287486528654 I[ 194 ]= 0.3432348010728626 R[ 194 ]= 766.5532419447005 F[ 194 ]= 153.3106483889401 Time[ 195 ]= 24.375 S[ 195 ]= 78.78687876798759 I[ 195 ]= 0.32348828829134074 R[ 195 ]= 766.5746941197675 F[ 195 ]= 153.31493882395353 Time[ 196 ]= 24.5 S[ 196 ]= 78.78122803129733 I[ 196 ]= 0.3048774033597461 R[ 196 ]= 766.5949121377857 F[ 196 ]= 153.31898242755716 Time[ 197 ]= 24.625 S[ 197 ]= 78.77590274853632 I[ 197 ]= 0.2873368808687715 R[ 197 ]= 766.6139669754957 F[ 197 ]= 153.32279339509915 Time[ 198 ]= 24.75 S[ 198 ]= 78.77088416209904 I[ 198 ]= 0.27080520124089447 R[ 198 ]= 766.63192553055 F[ 198 ]= 153.32638510611002 Time[ 199 ]= 24.875 S[ 199 ]= 78.76615459694194 I[ 199 ]= 0.2552243763049145 R[ 199 ]= 766.6488508556275 F[ 199 ]= 153.32977017112555 Time[ 200 ]= 25.0 S[ 200 ]= 78.761697397942 I[ 200 ]= 0.24053974708199083 R[ 200 ]= 766.6648023791466 F[ 200 ]= 153.33296047582937 Time[ 201 ]= 25.125 S[ 201 ]= 78.75749687089802 I[ 201 ]= 0.2266997930948191 R[ 201 ]= 766.6798361133392 F[ 201 ]= 153.3359672226679 Time[ 202 ]= 25.25 S[ 202 ]= 78.75353822696114 I[ 202 ]= 0.21365595254959002 R[ 202 ]= 766.6940048504076 F[ 202 ]= 153.3388009700816 Time[ 203 ]= 25.375 S[ 203 ]= 78.74980753029314 I[ 203 ]= 0.20136245277636716 R[ 203 ]= 766.707358347442 F[ 203 ]= 153.34147166948847 Time[ 204 ]= 25.5 S[ 204 ]= 78.74629164876367 I[ 204 ]= 0.18977615034760692 R[ 204 ]= 766.7199435007406 F[ 204 ]= 153.34398870014817 Time[ 205 ]= 25.625 S[ 205 ]= 78.7429782075084 I[ 205 ]= 0.17885638032680853 R[ 205 ]= 766.7318045101373 F[ 205 ]= 153.34636090202753 Time[ 206 ]= 25.75 S[ 206 ]= 78.73985554518089 I[ 206 ]= 0.16856481412981644 R[ 206 ]= 766.7429830339078 F[ 206 ]= 153.34859660678163 Time[ 207 ]= 25.875 S[ 207 ]= 78.73691267274079 I[ 207 ]= 0.1588653255101877 R[ 207 ]= 766.7535183347909 F[ 207 ]= 153.35070366695825 Time[ 208 ]= 26.0 S[ 208 ]= 78.73413923463035 I[ 208 ]= 0.14972386420736342 R[ 208 ]= 766.7634474176352 F[ 208 ]= 153.35268948352714 Time[ 209 ]= 26.125 S[ 209 ]= 78.73152547219993 I[ 209 ]= 0.14110833682222595 R[ 209 ]= 766.7728051591482 F[ 209 ]= 153.35456103182975 Time[ 210 ]= 26.25 S[ 210 ]= 78.72906218925144 I[ 210 ]= 0.13298849450905775 R[ 210 ]= 766.7816244301996 F[ 210 ]= 153.35632488604003 Time[ 211 ]= 26.375 S[ 211 ]= 78.7267407195763 I[ 211 ]= 0.12533582709601543 R[ 211 ]= 766.7899362111065 F[ 211 ]= 153.35798724222138 Time[ 212 ]= 26.5 S[ 212 ]= 78.72455289637205 I[ 212 ]= 0.11812346326806299 R[ 212 ]= 766.7977697002999 F[ 212 ]= 153.3595539400601 Time[ 213 ]= 26.625 S[ 213 ]= 78.72249102342806 I[ 213 ]= 0.11132607646693746 R[ 213 ]= 766.8051524167541 F[ 213 ]= 153.36103048335096 Time[ 214 ]= 26.75 S[ 214 ]= 78.72054784797777 I[ 214 ]= 0.10491979618221109 R[ 214 ]= 766.8121102965333 F[ 214 ]= 153.3624220593068 Time[ 215 ]= 26.875 S[ 215 ]= 78.7187165351204 I[ 215 ]= 0.09888212432592687 R[ 215 ]= 766.8186677837947 F[ 215 ]= 153.36373355675906 Time[ 216 ]= 27.0 S[ 216 ]= 78.7169906437212 I[ 216 ]= 0.09319185640067598 R[ 216 ]= 766.8248479165651 F[ 216 ]= 153.36496958331313 Time[ 217 ]= 27.125 S[ 217 ]= 78.71536410370442 I[ 217 ]= 0.08782900718741043 R[ 217 ]= 766.8306724075901 F[ 217 ]= 153.36613448151815 Time[ 218 ]= 27.25 S[ 218 ]= 78.71383119465797 I[ 218 ]= 0.08277474069479486 R[ 218 ]= 766.8361617205393 F[ 218 ]= 153.36723234410798 Time[ 219 ]= 27.375 S[ 219 ]= 78.7123865256741 I[ 219 ]= 0.07801130412654558 R[ 219 ]= 766.8413351418327 F[ 219 ]= 153.36826702836666 Time[ 220 ]= 27.5 S[ 220 ]= 78.71102501635413 I[ 220 ]= 0.07352196563703128 R[ 220 ]= 766.8462108483407 F[ 220 ]= 153.36924216966824 Time[ 221 ]= 27.625 S[ 221 ]= 78.70974187890992 I[ 221 ]= 0.06929095565846093 R[ 221 ]= 766.8508059711929 F[ 221 ]= 153.3701611942387 Time[ 222 ]= 27.75 S[ 222 ]= 78.7085326012987 I[ 222 ]= 0.06530341159530398 R[ 222 ]= 766.8551366559216 F[ 222 ]= 153.37102733118442 Time[ 223 ]= 27.875 S[ 223 ]= 78.70739293133114 I[ 223 ]= 0.061545325693215205 R[ 223 ]= 766.8592181191462 F[ 223 ]= 153.37184362382936 Time[ 224 ]= 28.0 S[ 224 ]= 78.70631886169666 I[ 224 ]= 0.05800349590070915 R[ 224 ]= 766.863064702002 F[ 224 ]= 153.37261294040053 Time[ 225 ]= 28.125 S[ 225 ]= 78.70530661585263 I[ 225 ]= 0.05466547955218438 R[ 225 ]= 766.8666899204958 F[ 225 ]= 153.3733379840993 Time[ 226 ]= 28.25 S[ 226 ]= 78.70435263472774 I[ 226 ]= 0.05151954971066804 R[ 226 ]= 766.8701065129678 F[ 226 ]= 153.3740213025937 Time[ 227 ]= 28.375 S[ 227 ]= 78.70345356419224 I[ 227 ]= 0.04855465401787011 R[ 227 ]= 766.8733264848247 F[ 227 ]= 153.37466529696508 Time[ 228 ]= 28.5 S[ 228 ]= 78.70260624325094 I[ 228 ]= 0.04576037590783424 R[ 228 ]= 766.8763611507009 F[ 228 ]= 153.3752722301403 Time[ 229 ]= 28.625 S[ 229 ]= 78.70180769291701 I[ 229 ]= 0.043126898048677476 R[ 229 ]= 766.8792211741951 F[ 229 ]= 153.37584423483915 Time[ 230 ]= 28.75 S[ 230 ]= 78.70105510572739 I[ 230 ]= 0.04064496788465138 R[ 230 ]= 766.8819166053231 F[ 230 ]= 153.37638332106476 Time[ 231 ]= 28.875 S[ 231 ]= 78.70034583586263 I[ 231 ]= 0.03830586515805851 R[ 231 ]= 766.8844569158159 F[ 231 ]= 153.37689138316333 Time[ 232 ]= 29.0 S[ 232 ]= 78.6996773898364 I[ 232 ]= 0.03610137129744539 R[ 232 ]= 766.8868510323883 F[ 232 ]= 153.3773702064778 Time[ 233 ]= 29.125 S[ 233 ]= 78.69904741772154 I[ 233 ]= 0.03402374056498863 R[ 233 ]= 766.8891073680944 F[ 233 ]= 153.377821473619 Time[ 234 ]= 29.25 S[ 234 ]= 78.69845370488204 I[ 234 ]= 0.032065672862117714 R[ 234 ]= 766.8912338518797 F[ 234 ]= 153.37824677037608 Time[ 235 ]= 29.375 S[ 235 ]= 78.6978941641813 I[ 235 ]= 0.030220288098195658 R[ 235 ]= 766.8932379564336 F[ 235 ]= 153.37864759128686 Time[ 236 ]= 29.5 S[ 236 ]= 78.69736682863962 I[ 236 ]= 0.02848110203252803 R[ 236 ]= 766.8951267244397 F[ 236 ]= 153.37902534488808 Time[ 237 ]= 29.625 S[ 237 ]= 78.6968698445146 I[ 237 ]= 0.026842003505109378 R[ 237 ]= 766.8969067933167 F[ 237 ]= 153.3793813586635 Time[ 238 ]= 29.75 S[ 238 ]= 78.69640146478046 I[ 238 ]= 0.02529723297636191 R[ 238 ]= 766.8985844185357 F[ 238 ]= 153.37971688370732 Time[ 239 ]= 29.875 S[ 239 ]= 78.6959600429829 I[ 239 ]= 0.023841362300690727 R[ 239 ]= 766.9001654955968 F[ 239 ]= 153.38003309911952 Time[ 240 ]= 30.0 S[ 240 ]= 78.69554402744805 I[ 240 ]= 0.022469275662988522 R[ 240 ]= 766.9016555807406 F[ 240 ]= 153.38033111614828
Just looking at numbers is boring.
Plotting the Results!¶
Let's Plot it! Here we are going to pull each value in sol.T so we can specifically customize each color.
################################################
#
# Plot Results
#
plt.plot(Time, S, "g", # susceptables
Time, I, "r", # infected
Time, R, "b", # recovered
Time, F, "k") # fatalities
plt.title("SIRF Model System, $R_o$="+
str(Ro) +
", $p_c$=" +
str(herd_immunity) +
", $p$=" +
str(immunization_fraction))
plt.xlim(0, TimePeriod)
plt.ylim(0, (S_0 + I_0 + R_0 + F_0))
plt.xlabel('Time (Days)')
plt.ylabel("Populations")
plt.legend(["Susceptables",
"Infected",
"Recovered",
"Fatalities"])
plt.show()
#
################################################
Exploring Herd Immunity¶
Let's now explore three values of Herd Immunity. We will run three cases with 25%, 50% and 75% of the populus being vaccinated.
################################################
#
# Plot Results
#
# We're also going to go for a longer game (one year)
TimePeriod = 365.
nTime = int(TimePeriod / DeltaT + 1)
Time = np.linspace(start = 0,
stop = TimePeriod, # this is *inclusive*
num = nTime)
#
# Setting up our arrays
#
S = Time*0 # Uninfected People (multiplying time by zero)
I = Time*0 # Infected (multiplying time by zero)
R = Time*0 # Recovered (multiplying time by zero)
F = Time*0 # Fatalities (multiplying time by zero)
#
# And we can initialize our first (or zero'th since it's python)
# time step from our above initial conditions
#
k = 0
S[k] = S_0 # Uninfected People (includes Vaccinated)
I[k] = I_0 # Infected
R[k] = R_0 # Recovered
F[k] = F_0 # Recovered
# This is the only time we need to do this above initialization
# until we change the size of our arrays.
# remember X[k=0] doesn't change in our loops
for immunization_fraction in [0.00,0.25, 0.50, 0.75, 1.00]:
############################################
#
# Run the Solver
#
for k in np.arange(start = 1, stop = nTime):
S[k] = S[k-1] + dSdt(S[k-1], I[k-1], R[k-1]) * DeltaT
I[k] = I[k-1] + dIdt(S[k-1], I[k-1], R[k-1]) * DeltaT
R[k] = R[k-1] + dRdt(I[k-1]) * DeltaT
F[k] = F[k-1] + dFdt(I[k-1]) * DeltaT
#
# For this graph we will also want to sum up the total
# number of people who have contracted the disease
#
Total_Impacted_Cases = I + R + F
#
# Plot Each Graph
#
plt.plot(Time, S, "g", # susceptables
Time, I, "r", # infected
Time, R, "b", # recovered
Time, F, "k", # fatalities
Time, Total_Impacted_Cases, "m") # impacted cases
plt.xlim(0, TimePeriod)
plt.ylim(0,(S_0 + I_0 + R_0 + F_0))
plt.title("SIRF Model System, $R_o$="+
str(Ro) +
", $p_c$=" +
str(herd_immunity) +
", $p$=" +
str(immunization_fraction))
plt.xlabel('Time (Days)')
plt.ylabel("Populations")
plt.legend(["Susceptables",
"Infected",
"Recovered",
"Fatalities",
"Impacted"])
plt.show()
#
############################################
#
################################################
Since this is more about learning programming tricks in Python rather than learning we can do this again... not with a loop but with a 4-panel.
I'm going to "steal" the 4-panel section from our plotting tutorial for a lot of this material because I forget this operation way too often.
#############################################
#
# Make a 2x2 plot with differennt logs on the
# axes to demonstrate leveraging the ax-and-fig
# method of plotting.
#
#
# We'll start with the subplot command which
# will let us make 2 rows (first argument)
# and 2 columns.
#
# The second modifier pair instruct the matplotlib
# "share" the axis attributes for each plot.
#
# When we do this, FIG will represent the atributes
# of the whole plotting area.
#
# The acutal "subplots" are typically called "ax"
# which you can consider to be the
#
fig, ax = plt.subplots(2, # 2 rows in the y
2, # 2 cols in the x
sharex='col', # set the colum (x axes to match)
sharey='row') # set the rows (y axes to match)
# # # # # # # # # # # # # # # # # # # # # # # # #
# Let's have our Top Left (0,0) plot represent the p=0 case
# # # # # # # # # # # # # # # # # # # # # # # # #
immunization_fraction = 0.00
for k in np.arange(start = 1, stop = nTime):
S[k] = S[k-1] + dSdt(S[k-1], I[k-1], R[k-1]) * DeltaT
I[k] = I[k-1] + dIdt(S[k-1], I[k-1], R[k-1]) * DeltaT
R[k] = R[k-1] + dRdt(I[k-1]) * DeltaT
F[k] = F[k-1] + dFdt(I[k-1]) * DeltaT
#
# For this graph we will also want to sum up the total
# number of people who have contracted the disease
#
Total_Impacted_Cases = I + R + F
ax[0, 0].plot(Time, S, "g", # susceptables
Time, I, "r", # infected
Time, R, "b", # recovered
Time, F, "k", # fatalities
Time, Total_Impacted_Cases, "m") # impacted cases
ax[0, 0].set_title("$p$="+str(immunization_fraction),
loc='left')
# Let's have our Top Right (0, 1) plot represent the p=0.25 case
# we are only going to put the legend on one pannel
# (this Lower Right one)
immunization_fraction = 0.25
for k in np.arange(start = 1, stop = nTime):
S[k] = S[k-1] + dSdt(S[k-1], I[k-1], R[k-1]) * DeltaT
I[k] = I[k-1] + dIdt(S[k-1], I[k-1], R[k-1]) * DeltaT
R[k] = R[k-1] + dRdt(I[k-1]) * DeltaT
F[k] = F[k-1] + dFdt(I[k-1]) * DeltaT
#
# For this graph we will also want to sum up the total
# number of people who have contracted the disease
#
Total_Impacted_Cases = I + R + F
ax[0, 1].plot(Time, S, "g", # susceptables
Time, I, "r", # infected
Time, R, "b", # recovered
Time, F, "k", # fatalities
Time, Total_Impacted_Cases, "m") # impacted cases
ax[0, 1].set_title("$p$="+str(immunization_fraction),
loc='right')
# # # # # # # # # # # # # # # # # # # # # # # # #
# Let's have our Lower Left (1, 0) plot represent the p=0.25 case
# # # # # # # # # # # # # # # # # # # # # # # # #
immunization_fraction = 0.50
for k in np.arange(start = 1, stop = nTime):
S[k] = S[k-1] + dSdt(S[k-1], I[k-1], R[k-1]) * DeltaT
I[k] = I[k-1] + dIdt(S[k-1], I[k-1], R[k-1]) * DeltaT
R[k] = R[k-1] + dRdt(I[k-1]) * DeltaT
F[k] = F[k-1] + dFdt(I[k-1]) * DeltaT
#
# For this graph we will also want to sum up the total
# number of people who have contracted the disease
#
Total_Impacted_Cases = I + R + F
ax[1, 0].plot(Time, S, "g", # susceptables
Time, I, "r", # infected
Time, R, "b", # recovered
Time, F, "k", # fatalities
Time, Total_Impacted_Cases, "m") # impacted cases
ax[1, 0].set_title("$p$="+str(immunization_fraction),
loc='left')
# # # # # # # # # # # # # # # # # # # # # # # # #
# Let's have our Lower Right (1, 1) plot represent the p=0.75 case
# which for us is on the safer side of our needed herd immunity
# the legend goes here because it's rather big.
# # # # # # # # # # # # # # # # # # # # # # # # #
immunization_fraction = 0.75
for k in np.arange(start = 1, stop = nTime):
S[k] = S[k-1] + dSdt(S[k-1], I[k-1], R[k-1]) * DeltaT
I[k] = I[k-1] + dIdt(S[k-1], I[k-1], R[k-1]) * DeltaT
R[k] = R[k-1] + dRdt(I[k-1]) * DeltaT
F[k] = F[k-1] + dFdt(I[k-1]) * DeltaT
#
# For this graph we will also want to sum up the total
# number of people who have contracted the disease
#
Total_Impacted_Cases = I + R + F
ax[1, 1].plot(Time, S, "g", # susceptables
Time, I, "r", # infected
Time, R, "b", # recovered
Time, F, "k", # fatalities
Time, Total_Impacted_Cases, "m") # impacted cases
ax[1, 1].set_title("$p$="+str(immunization_fraction),
loc='right')
ax[1, 1].legend(["Susceptables",
"Infected",
"Recovered",
"Fatalities",
"Total Impacted"])
# add the big title
fig.suptitle("SIRF Model System, $R_o$="+
str(Ro) +
", $p_c$=" +
str(herd_immunity))
# you have to kludge a common axis here which is
# rather disapointing (maybe the next version of
# matplotlib will have it.)
#
# we use the fig.text function
# the first two arguments are the x and y centers of
# the plot in the larger plotter space
#
fig.text(0.50, 0.00,
'Time (days)',
ha = 'center')
fig.text(0.00, 0.50,
'Populations',
va = 'center',
rotation = 'vertical')
plt.show()
#
############################################
Exploring Herd Immunity¶
Let's look more closely at how the herd immunity chagnes the overall impact of the diseases (counting results)
This also shows how to loop through the system and load the results into an array
################################################
#
# Plot Results
#
#
# let's first make an array to carry our
# immune factor
#
# Make the immunity (dp = 0.01 so we have 101 slots)
immune_array = np.linspace(0, 1,101)
# clone the immune array as zeros for the impact array
impact_array = np.zeros(immune_array.size)
for i in range(0, immune_array.size):
####################################
#
# Load Immunization Fraction
#
immunization_fraction = immune_array[i]
#
# Run the ODE Solver
#
for k in np.arange(start = 1, stop = nTime):
S[k] = S[k-1] + dSdt(S[k-1], I[k-1], R[k-1]) * DeltaT
I[k] = I[k-1] + dIdt(S[k-1], I[k-1], R[k-1]) * DeltaT
R[k] = R[k-1] + dRdt(I[k-1]) * DeltaT
F[k] = F[k-1] + dFdt(I[k-1]) * DeltaT
#
# For this graph we will also want to sum up the total
# number of people who have contracted the disease
#
Total_Impacted_Cases = I + R + F
#
#
impact_array[i] = Total_Impacted_Cases[nTime-1]
#
####################################
plt.plot(immune_array,
impact_array,
"m")
plt.vlines(herd_immunity,
0, (S_0 + I_0 + R_0 + F_0),
'b')
plt.legend(["Total Impacted",
"Herd Immunity"])
plt.xlim(0, 1)
plt.ylim(0, (S_0 + I_0 + R_0 + F_0))
plt.title("SIRF Model System, $R_o$="+
str(Ro) +
", $p_c$=" +
str(herd_immunity))
plt.xlabel("Fraction of Population Immunized, $p$")
plt.ylabel("Total Impacted by Disease")
plt.show()
#
################################################
So if you don't have a legitimate medical reason not to take a given immunization (e.g., alergies to the flu vaccines, immunocompromised, etc.), GET YOUR FRIGGIN' SHOTS!!!!!
(and if your curious about the title of the exercise... click below.)
#Tom Lehrer Sing's "I got it from Agness"
YouTubeVideo('R6qFG0uop9k')
Version Information¶
################################################################
#
# Loading Version Information
#
%load_ext version_information
%version_information version_information, numpy, scipy, matplotlib
#
################################################################
| Software | Version |
|---|---|
| Python | 3.12.9 64bit [Clang 18.1.8 ] |
| IPython | 9.1.0 |
| OS | macOS 15.4 arm64 arm 64bit |
| version_information | 1.0.4 |
| numpy | 2.2.4 |
| scipy | 1.15.2 |
| matplotlib | 3.10.1 |
| Mon Apr 14 15:51:59 2025 MDT | |
